Research Approach
Our research group investigates how environmental exposures, biological susceptibility, and social context interact to shape health across populations and regions. Because humans and wildlife experience complex, location-specific mixtures of chemical and non-chemical stressors, our work integrates geospatial exposure science, mechanistic toxicology, and systems biology to understand how real-world environments contribute to disease.
Through this framework, we aim to advance precision environmental health by improving how environmental risk is measured, interpreted, and prevented. Our work emphasizes open science, reproducibility, and translational impact, supported by open-source computational tools and publicly accessible data and code.
Our research focuses on three interconnected areas:
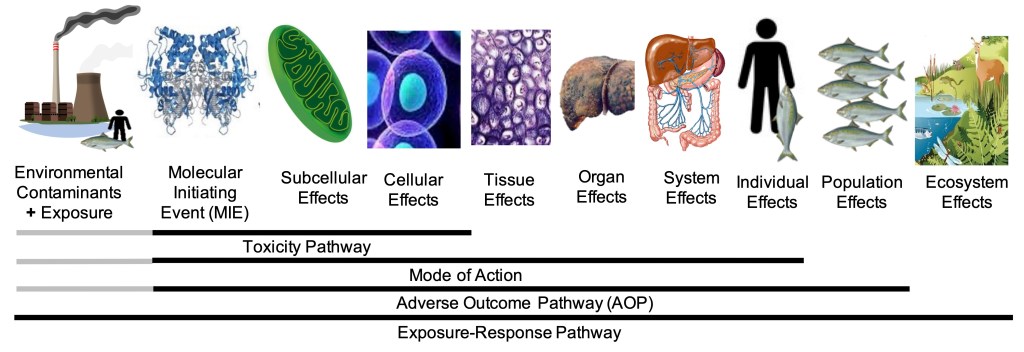
- Geospatial chemical mixtures and health: We quantify how spatially heterogeneous mixture exposures influence biological responses and disease risk using new approach methods, high-throughput phenotypic profiling, in vivo anchoring, and computational mixtures modeling linked to adverse outcome pathways.
- Interindividual susceptibility: We investigate how genetic and metabolic variability, particularly in cytochrome P450 enzymes, alters internal dose and toxicological response through engineered cell systems, toxicokinetic modeling, and population-level spatial data.
- Cumulative environmental and social stressors: We integrate chemical exposures with social vulnerability and structural determinants of health to identify regions where combined stressors disproportionately elevate disease risk and to inform place-based prevention strategies.
Current Research
Research Scientist – Computational Toxicology Research Group, Health Canada
Physiological modeling of immune responses to PFAS
Per- and polyfluoroalkyl substances (PFAS) are widespread environmental contaminants associated with immune dysfunction and other adverse health outcomes. Despite growing epidemiological and experimental evidence, key gaps remain in understanding how external exposure translates to internal dose, immune perturbation, and ultimately human health risk. This work applies a translational toxicology framework grounded in the 3Rs (Replacement, Reduction, and Refinement) by integrating complementary in vivo, in vitro, and in silico approaches.
- In vivo studies characterize immune cell populations, immune function, and tissue-specific molecular responses across relevant exposure levels while minimizing animal use.
- In vitro assays in immune and hepatic cell systems provide mechanistic insight and enable higher-throughput, human-relevant testing.
- In silico modeling, including physiologically based toxicokinetic (PBTK) and immune-response models, links external exposure to internal target-site dose and downstream biological effects.
Together, these data streams establish a quantitative, ethical, and mechanistically informed framework for evaluating PFAS immunotoxicity and strengthening translation to human health risk assessment.
Publications:
- PFOS and PFOA exposure induces liver injury and sex-dependent immune effects in C57BL/6 mice (Blais and Loan et al., 2026)
- A Novel Phase Extraction (SPE) Method to Quantify 21 Per-and Polyfluoroalkyl Compounds in Biological Samples (Guo et al., 2026)
Advancing risk assessment of complex chemical mixtures
This project examines the environmental and human health implications of bitumen extraction and processing, with a focus on naphthenic acids, metals, and polycyclic aromatic compounds. It integrates experimental characterization of exposure pathways and uptake with molecular and cellular toxicology approaches to assess liver toxicity and underlying mechanisms of action within a whole mixtures assessment framework. An integrated case-study risk assessment combines new experimental findings with existing environmental evidence to provide a comprehensive evaluation of complex environmental mixtures. The resulting insights are intended to inform regulatory decision-making, exposure guidelines, and risk-management strategies.
High-throughput phenotypic profiling for chemical safety assessment
This work advances Cell Painting and computational modeling capabilities to improve the prediction of chemical effects on biological systems and to support modernized chemical risk assessment.Two complementary goals guide this research:
- Regulatory translation: Applying these tools to evaluate chemical safety, compare cell models and profiling platforms, and demonstrate their value for hazard identification and decision-making while reducing reliance on animal testing.
- Method development: Refining high-throughput phenotypic profiling to generate reproducible, information-rich measures of biological response across different cellular models.
Publications:
- Smaller Scale, Same Impact: Replicating High-Throughput Phenotypic Profiling in a Medium-Throughput Lab for Use in Chemical Risk Assessment (Cho et al., 2025)
Previous Research
Postdoctoral Research
Division of Translational Toxicology (DTT), National Institute of Environmental health Sciences (NIEHS)
At NIEHS, my research examined how environmental chemical mixtures influence human health and how geospatial data and modeling can improve exposure-risk characterization in large population studies.
Publications:
- Lessons learned from evaluating defined chemical mixtures in a high-throughput estrogen receptor assay system (Parham et al., 2025)
- A geospatial modeling approach to quantifying the risk of exposure to environmental chemical mixtures via a common molecular target (Eccles et al., 2023)
- Integrating Multiscale Geospatial Environmental Data into Large Population Health Studies: Challenges and Opportunities (Cui et al., 2022)
- Air pollutant mixtures and reported psoriasis or eczema in the Personalized Environment and Genes Study (PEGS) (Lowe et al., 2023)
University of Toronto Mississauga
This research applied paleoecotoxicological approaches, including tree rings and sediment cores, to reconstruct historical contaminant baselines and evaluate long-term biological effects of environmental contamination. These methods enable assessment of regulatory effectiveness, reconstruction of historical exposure trends, and improved risk assessment for both wildlife and human populations.
Publications:
- Using sediment cores to reconstruct mining related deposition in Yellowknife, NWT, Canada (Cheney et al., 2020 and Cheney et al., 2024)
- Paleotoxicity of petrogenic and pyrogenic hydrocarbon mixtures in sediment cores (Thomas et al., 2022)
- Using tree-rings to reconstruct historical Hg concentrations in Yukon, Canada (Eccles et al., 2020)
Ph.D. Dissertation (University of Ottawa)
My doctoral work advanced ecological and human risk assessment by integrating environmental monitoring, biomarker development, and geospatial analysis to quantify spatial patterns of contaminant exposure and biological response. Key contributions included:
1. Developing a geospatial framework to improve ecological risk assessment:
- The Use of Geographic Information Systems for Spatial Ecological Risk Assessments (Eccles et al., 2019)
2. Developing fur as a non-invasive biomarker medium:
- Meta-regressions to derive conversion factors between Hg tissue concentration (Eccles et al., 2017)
- Quantifying the distribution of Hg across a pelt (Eccles et al., 2019)
3. Quantifying spatial patterns of exposure and effects using biomonitoring data:
- Spatial patterns of the exposure-response relationship between mercury and cortisol in the fur of river otter (Lontra canadensis) (Eccles et al., 2021)
- Geospatial analysis of the patterns of chemical exposures among biota in the Canadian Oil Sands Region (Eccles et al., 2020)
- Determining chemical sources from biomonitoring data (Eccles et al., 2020)